SCHUNTER LAB Molecular Ecology & Evolution
Molecular Ecology & Evolution
Ecological genomics, transcriptomics & epigenomics of adaptation, response to environmental change, transgenerational inheritance, and environmental effects on behaviour and the brain in marine organisms

The sea, once it casts its spell, holds one in its net of wonder forever – Jacques Yves Cousteau
RESEARCH
Effects of Climate Change
How do organisms deal with the rapid change in the environment? What is the effect of climate change on marine organisms?
How do organisms already live in e.g. acidic environments?

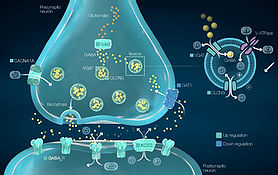
Behaviour & the Brain / Neurogenomics
Behaviour is often altered with a change in the surrounding environment. We want to understand how the environment can influence behaviour and how neuro-molecular states in the brain can drive this behavioural change.
Transgenerational & Parental Effects
Organisms react to their environment. Do they then pass on what they have learned to the next generation? If so, how?
We want to understand the mechanisms of parental effects and if they can help adaptation to future climate change conditions.

Population Connectivity & Ecological Genomics
How populations are connected or isolated is important in terms of local adaptation as well as population resilience. We study how recruitment of larvae to marine populations allows for population connectivity and investigate how isolated populations survive.

PEOPLE
PRINCIPAL INVESTIGATOR
Celia Schunter
Assistant Professor
Swire Institute of Marine Science
School of Biological Science
The University of Hong Kong
schunter(at)hku.hk

POSTDOCTORAL FELLOWS
Dr. Natalia Petit
is passionate about marine conservation and is interested in assessing the adaptive potential to global change in marine species
Dr. Kang Jingliang
is from China. He obtained his Bachelor’s degree in Aquaculture from Southwest University in 2013 and received his Ph.D in Aquatic Biology from the Chinese Academy of Sciences in 2018. He joined our team as a postdoc in August 2019. During his Ph. D, he focused on the high-altitude adaptation analysis of Creteuchiloglanis macropterus by population genetics (RAD-seq) and RNA-seq, which is a glyptosternoid fish that inhabits in Yunnan Province and Tibet, China. Kang is interested in investigating the molecular responses of fishes to specific environments through genome scanning.
Currently, he is investigating the molecular response of coral reef fishes to living in natural CO2 seeps (lower pH) by comparing the differences in brain gene expression in fish from a CO2 seep to nearby control reefs. He aims to detect the putative genes that are related to control of the cellular responses to elevated CO2 and to provide the common but also variable molecular mechanisms among species in coping with elevated CO2.
In his free time, Kang plays basketball, swims and likes watching sports.
Ph.D STUDENTS
Sneha Suresh
is originally from India and received her Bachelor’s degree in biotechnology from Ramaiah Institute of Technology in 2017 and a Master’s in Biology from the University of Massachusetts Lowell, USA, in 2019. Her Master’s thesis was focused on investigating the effects of copper toxicity on alternative splicing in Daphnia. Sneha is fascinated by how organisms regulate their genome to deal with changes in the environment.
As part of her Ph.D. thesis, she is investigating the gene expression responses in the spiny damselfish to conditions of ocean acidification. Her aim is to understand the influence of parental exposure on the offsprings’ response to pH change. While changes in ocean chemistry can have short-term negative effects on fishes and other marine organisms, they have the potential to acclimate to the acidic environments across generations. Intergenerational acclimation could, therefore, have long-lasting effects on future generations enabling them to adjust to more acidic environments. During her Ph.D. Sneha hopes to better understand the transcriptomic and epigenetic basis of intergenerational acclimation. She mainly relies on bioinformatics tools and approaches in her research to analyze and interpret an organism’s response to environmental change at the molecular level.
While not in the lab she enjoys hiking, reading mystery/ adventure fiction, scuba-diving and playing the violin.
Jade Sourisse
is from France, she studied at Lille University where she received her Bachelor’s in Life Sciences in 2017 and she graduated from Sorbonne University in Paris with a Master’s degree in Biodiversity and Conservation of Croisieres Marines. For her Master’s thesis, she studied the impact of habitat quality on gut microbiome communities in the foureye butterflyfish Chaetodon capistratus.
Jade is interested in using the power of molecular studies to investigate the effect of climate change and human activities on marine organisms and ecosystems.
For her Ph.D. thesis, Jade will focus on the effects of ocean acidification on the behaviour and related learning abilities and gene expression of the California sea hare (Aplysia californica). Unlike other marine species, the nervous system of Aplysia californica is well-mapped and neurons involved in specific behaviour have been identified.
Jade’s aim is to describe the behavioural effects of environmental change and observe the underlying gene expression changes in the involved components of the nervous system with ocean acidification.
Jade has also been a scuba diver since she was 8 years old. She learned to dive in the North of France and has traveled in many places since in order to achieve one of her life goals: dive in all the seas in the world!
MPHIL STUDENTS
Sandra Ramirez
is originally from Cali, Colombia. She studied in Universidad del Valle, where she obtained her degree in Biology with an emphasis on Marine studies in 2016. Her thesis was on the population genetics and connectivity of a damselfish (Stegastes flavilatus) among marine protected areas in the Colombian Pacific. After that, Sandra participated in a variety of studies involving fish ecology and coral plasticity in her coral reef research group and lead projects about local ecosystem conservation in Cali to finally join the Schunter lab in April 2019.
Currently, Sandra’s research involves the study of the molecular mechanisms underlying the cleaning behaviour of the coral reef fish Labroides dimidiatus. She is interested in identifying the molecular pathways that are associated with the interactions of cleaner fish and its potential clients, and how different environmental changes such as near-future ocean acidification and warming conditions may affect them as well. In addition, Sandra is interested in analyzing potential evolutionary adaptations that these organisms may have when exposed to different environmental conditions.
Even though Sandra loves diving and exploring the sea, she also loves to go into the jungle and observe terrestrial animals such as birds, frogs, and mammals. She loves to go into fieldwork but enjoys working in the lab as well. She is also a fan of salsa music, movies, and travel!
Arthur Chung
is from Hong Kong and received his Bachelor’s degree in Ecology & Biodiversity from the University of Hong Kong (HKU) in 2017. As a research assistant after his studies, he worked on applying population genomic data into marine restoration of local gorgonians. Such experience had made him recognise his passion in pursuing marine conversation by utilising the power of molecular tools and techniques.
As part of his MPhil thesis, he is investigating the ecological impacts brought from the mercy release of hybrid grouper (Epinephelus fuscoguttatus x E. lanceolatus) into the wild by performing gut content analysis with metabarcoding. The hybrid grouper is carnivorous and released into the wild in large quantities, yet there is a complete lack of information on the ecology of the wild populations.
His ultimate goal is to utilize genomic data to understand the speciation processes within the genus Epinephelus and reveal if any hybridization events occurred in the past that are contributing to species diversification.
RESEARCH STUDENTS
Iris Tan
Is interested in marine life and passionate about marine conservation. Loves nature and travel.
Currently working on using COI nucleotide diversity as an early diagnostic of the conservation status of commercial fishes from the South China Sea.
Stanley Chan
is interested in genetic, evolution, and different kind of strange creatures. Also loves reading, gaming, and drinking!
Currently working alternative splicing patterns underlying the response to a heatwave in a wild fish from the Great Barrier Reef.
STAFF
Jessie Lai
Technical Support & Magician!
is on the constant hunt for Northern Lights.
FORMER LAB MEMBERS
Rainbow Tsang
worked in the lab as a Research Assistant for a year. She is passionate about marine conservation and education
is working on proteomic signals underlying exposure to ocean acidification in a coral reef fish
Elliot Lee
is interested in neurosciences and marine life. Currently working on RNA alternative splicing events in fish brains.
loves outdoor activities and a good drinking night!
Max Tin
is interested in marine biology, especially fish. Loves nature and its wonders.
Currently working on alternative RNA splicing in Amphiprion percula under diel CO2 cycles (orange clownfish)
PUBLICATIONS
Tsang R, Welch M, Munday P, Ravasi T Schunter C* (2020) Proteomic responses to ocean acidification in the brain of juvenile coral reef fish. Frontiers in Marine Science. 10.3389/Fmars.2020.00605
Bernal M*, Schunter C*, Lehmann R, Lightfoot, DJ, Allan BJM, Veilleux HD, Rummer JL, Munday PL, Ravasi T (2020) Species-specific molecular responses of wild coral reef fishes during a marine heatwave. Science Advances. 10.1126/sciadv.aay3423 *contributed equally. https://advances.sciencemag.org/content/6/12/eaay3423
New coverage of the fish heatwave paper:
https://www.tarifs.org/croisiere-sur-la-seine
https://phys.org/news/2020-03-seas-climate-toll-tropical-fish.html
Monroe A, Zhang H, Schunter C*, Ravasi T* (2020) Probing SWATH-MS as a tool for proteome level quantification in a non-model fish. Molecular Ecology Resources. 10.1111/1755-0998.13229
Schunter C, Ravasi T, Munday PL, Nilsson G (2019) Neural effects of elevated CO2 in fish may be amplified by a vicious cycle. Conservation Physiology. 10.1093/conphys/coz100
Munday P, Schunter C, Allan B, Nicol S, Parsons D, Pether S, Ravasi T, Setiawan A, Smith N & Domingos J (2019) Testing the adaptive potential of yellowtail kingfish to ocean warming and acidification. Front. Ecol. Evol. 7: 253. doi: 10.3389/fevo
Schunter C, Macpherson E, Raventos N, Garriga J, Garza JC, Bartumeus F, Pascual M (2019) A novel integrative approach elucidates fine-scale dispersal patchiness in marine populations. Scientific Reports 9: 10796 (2019)

Special Issue on "The role of plasticity in phenotypic adaptation to rapid environmental change’
guest edited by Donelson JM, Fox RJ, Gaitan-Espitia JD, Schunter C and Ravasi T
Fox JR, Donelson JM, Schunter C, Ravasi T, Gaitan-Espitia JD (2019) Beyond buying time: the role of plasticity in phenotypic adaptation to rapid environmental change. https://doi.org/10.1098/rstb.2018.0174
Schunter C, Lehmann R, Bernal M, Ravasi T (2018) Unusual bilateral color pattern in a regal angelfish from the Red Sea. Bulletin of Marine Science. doi.org/10.5343/bms.2018.0050
Youtube video to the note: https://youtu.be/pGdaKYOvGyY
Lehmann R, Lightfoot D, Schunter C, Mitchell CT, Ohyanagi H, Mineta K, Foret S, Berumen M, Miller DJ, Aranda M, Gojobori T, Munday P, Ravasi T (2018) Finding Nemo’s Genes: A chromosome-scale reference assembly of the genome of the orange clownfish Amphiprion percula. Molecular Ecology Resources.doi.org/10.1111.1755-0998.12939
Schunter C, Bonzi L, Norstog J, Parkes S, McCabe M, Ravasi T (2017) Desert fish populations defy hydrological constraints via ecological and physiological adaptation. (under review).
Schunter C, Welch M, Nilsson G, Rummer J, Munday P, Ravasi T (2017) An interplay between plasticity and parental phenotype determines impacts of ocean acidification on a reef fish. Nature Ecology & Evolution. doi:10.1038/s42559/017-0428-8.
Boero F, Foglini F, Fraschetti S, Goriup P, Macpherson E, Planes S, Soukissian T, The CoCoNet Consortium (2017) CoCoNet: towards coast to coast networks of marine protected areas (from the shore to the high and deep sea), coupled with sea-based wind energy potential. SCIentific RESearch and Information Technology, 6, 1-95.
Pascual M, Rives B, Schunter C, Macpherson E (2017) Impact of life history traits on gene flow: A multispecies systematic review across oceanographic barriers in the Mediterranean Sea. PLoS One.
doi.org/10.1371/journal.pone.0176419
Schunter C, Welch M, Ryu T, Zhang H, Nilsson G, Munday PL, Ravasi T (2016) Molecular signatures of transgenerational response to ocean acidification in a species of reef fish. Nature Climate Change, 6, 1014-1018. doi:10.1038/ncimate3087. Altimetrics score (April 2017): 384.
Ryu T, Seridi L, Moitinho-Silva L, Oates M, Liew Y, Mavromatis C, Wang X, Haywood A, Lafi F, Kupresanin M, Sougrat R, Alzahrani M, Giles E, Ghosheh Y, Schunter C, Baumgarten S, Berumen M, Gao X, Aranda M, Foret S, Gough J, Voolstra CR, Hentschel U, Ravasi T (2016) Hologenome analysis of two marine sponges with different microbiomes. BMC Genomics, 17, 158.
Schunter C, Pascual M, Garza JC, Raventos N, Macpherson E (2014) Kinship analyses identify fish dispersal events on a temperate coast line. Proceedings of the Royal Society B, 281, 20140556.
Schunter C, Vollmer S, Macpherson E, Pascual M (2014) Transcriptome analyses and differential gene expression in a non-model fish species with alternative mating tactics. BMC Genomics, 15, 167.
Schunter C, Garza C, Macpherson E, Pascual M (2014) SNP development from RNA-seq data in a non-model fish: how many individuals are needed for accurate allele frequency prediction? Molecular Ecology Resources, 14, 157–165.
Schunter C, Carreras-Carbonell J, Macpherson E, Tintoré J, Vidal-Vijande E, Pascual A, Guidetti P, Pascual M (2011) Matching genetics with oceanography: directional gene flow in a Mediterranean fish species. Molecular Ecology, 20, 5167–81.
Schunter C, Carreras-Carbonell J, Planes S, Sala E, Ballesteros E, Zabala M, Harmelin J-G, Harmelin-Vivien M, Macpherson E, Pascual M (2011) Genetic connectivity pattern in an endangered species: the dusky grouper (Epinephelus marginatus). Journal of Experimental Marine Biology and Ecology, 401 (1-2), 126-133.
Gerlach G, Hotchkins-Davis A, Avolio C, Schunter C (2008) Kin recognition in Zebrafish: a 24-hour window for olfactory imprinting. Proceedings of the Royal Society: Biological Sciences. 275: 1647, 2165 -2170
SCHUNTER LAB
Swire Institute of Marine Science;
School of Biological Sciences,
Kadoorie Biological Sciences Building,
The University of Hong Kong,
Pok Fu Lam Road,
Hong Kong SAR
schunter(at)hku.hk